A deep dive into hybridization capture for targeted sequencing
Targeted NGS: A cost-effective approach to genomic research
Whole genome sequencing is powerful, but it isn’t always the right tool for the job. When your research questions center on specific genes, genomic regions, or curated panels of variants, targeted sequencing offers a smarter, more cost-effective approach. By focusing sequencing efforts on regions of interest rather than the entire genome, targeted NGS delivers deep, high-quality coverage precisely where it is needed, while eliminating irrelevant data from downstream bioinformatic analysis.
The result is a dramatically more efficient use of sequencing resources: lower per-sample costs, cleaner data, and faster time to insight. For this approach, many researchers will use either PCR-based amplicon sequencing or probe-based hybridization capture. The choice between these methods depends on the size of the target, the quality of the starting material, and the required uniformity of coverage. In this blog, we will compare these methods, walk through how hybridization capture works, and explore the research applications where it truly shines.
Hybridization capture vs. amplicon-based methods for targeted sequencing
When it comes to targeted sequencing, researchers typically choose between two approaches: PCR-based amplicon sequencing and probe-based hybridization capture. Each has its strengths, and understanding their differences helps you match the method to your specific research needs.
Amplicon sequencing (PCR-based)
Amplicon sequencing works by designing PCR primers that flank specific regions of interest and amplifying those targets directly from genomic DNA. It’s a fast, cost-effective approach particularly well-suited to small, custom NGS panels targeting a defined set of known variants. It is a method best reserved for detecting known SNVs and small indels in well-characterized regions. Because it requires relatively little input DNA, it can be suitable for low-input libraries. However, amplicon methods struggle with complex targets, large panels, and sequence-uniform regions where primer design becomes challenging.1,2
Hybridization capture (probe-based)
Hybridization capture takes a different approach. Rather than amplifying targets by PCR, it uses biotinylated oligonucleotide probes, or “baits”, designed to be complementary to the regions of interest. These probes hybridize to the targets in solution, and the resulting probe-target complexes are physically pulled down using streptavidin-coated magnetic beads, effectively enriching the library for the sequences you care about.3
The advantages are substantial. Hybridization capture delivers better on-target coverage uniformity across the panel, handles large and complex genomic regions with greater reliability, and excels at detecting novel rearrangements and structural variants that amplicon methods might miss entirely. It also performs well with challenging sample types, including formalin-fixed, paraffin-embedded (FFPE) tissue and other degraded or fragmented DNA sources, making it an indispensable tool for fields that routinely work with archival or ancient material. For researchers building comprehensive, multi-gene panels or tackling non-model organisms without well-characterized genomes, hybridization capture is the method of choice.1,2
The hybridization capture workflow
Following standard NGS library preparation, a hybridization capture experiment proceeds through three main steps (Figure 1):
- Hybridization: The prepared library is denatured into single strands and incubated with biotinylated oligonucleotide probes (baits) designed to be complementary to your targets of interest. During incubation, the probes bind specifically to their complementary target sequences in the library.3
- Capture and wash: Streptavidin-coated magnetic beads are added to bind the biotinylated probes. A magnet then separates the probe-target hybrids from the rest of the library, which is washed away, leaving behind only the enriched, on-target molecules.3
- Amplification: The captured, target-enriched library is released from the baits and amplified in preparation for sequencing.3
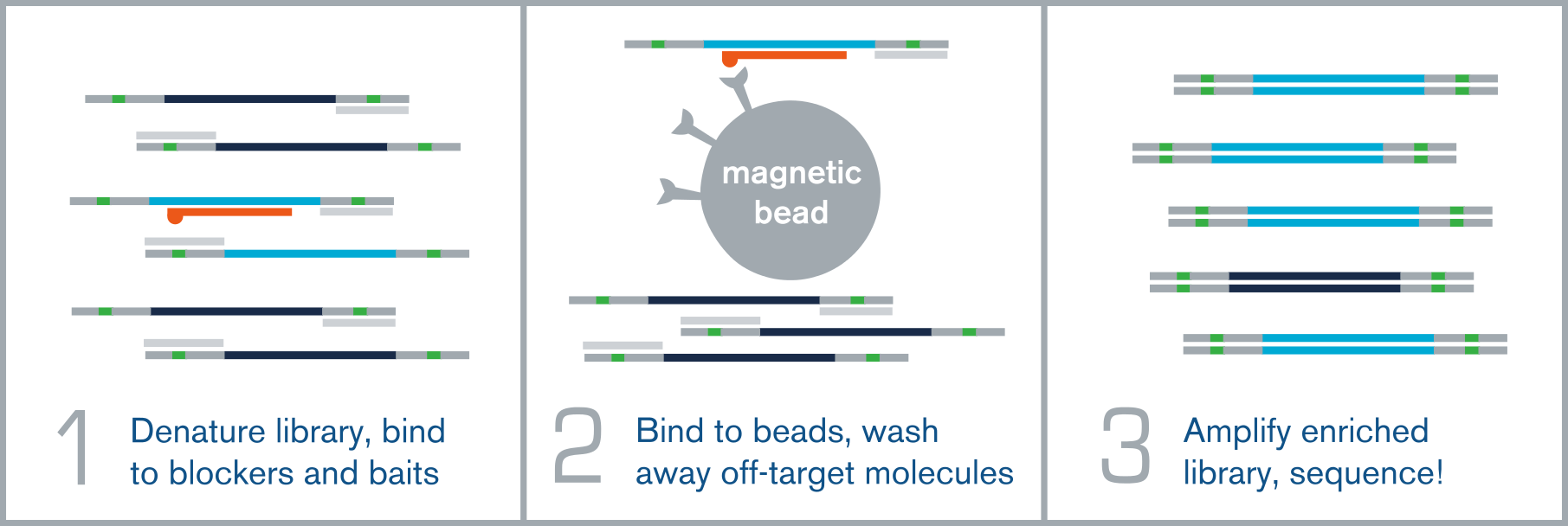
Figure 1. An illustration of the three main phases of the hybridization workflow.
Introducing myBaits® by Daicel Arbor Biosciences
Daicel Arbor Biosciences’ myBaits system is purpose-built to make hybridization capture accessible, flexible, and scalable for any research program. Using a proprietary oligo pool synthesis technology, myBaits kits deliver optimized chemistry and even coverage across a wide range of target types and organisms (human, plant, animal, bacterial, viral, ancient DNA, and more).
A key differentiator is the platform’s openness and flexibility:
- Compatible with any NGS library preparation system, so it integrates seamlessly into existing workflows.
- Scalable kit and panel sizes to fit projects from focused gene panels to genome-wide enrichment.
- Complete kits include all necessary hybridization and wash reagents, simplifying setup and reducing the risk of protocol variability.
Researchers can also design fully custom myBaits panels for maximum flexibility, tailoring probe sets to their exact targets of interest, whether that’s a curated set of SNPs in a livestock genome or a suite of pathogen resistance genes. Our scientists can assist with project scoping, panel design, and bait sequence optimization to reduce your per-sample costs while increasing the accuracy and power of your data analysis.
Applications where hybridization capture excels
The versatility of hybridization capture has made it a go-to approach across a remarkably broad range of research areas.
Cancer research
In cancer genomics, hybridization capture enables the construction of large, comprehensive oncology panels that simultaneously interrogate hundreds of genes for somatic variants, copy number alterations, and gene fusions. Its ability to handle FFPE samples, the standard format for archived tumor tissue, and detect novel rearrangements makes it especially well-suited for translational research and biomarker discovery.1
Agrigenomics
Agricultural researchers use targeted gene sequencing to efficiently screen for trait-associated markers across large breeding populations. Whether characterizing disease resistance in crops, genotyping livestock, or monitoring pathogen diversity in agricultural ecosystems, hybridization capture provides the uniform, deep coverage needed to make confident calls across complex, polyploid genomes.
Epigenetic sequencing
Methylation sequencing is a growing application where hybridization capture adds real value. By combining bisulfite conversion or enzymatic methylation profiling with targeted capture, researchers can interrogate methylation patterns at specific loci with much greater depth and efficiency than whole-genome methylation sequencing alone.
Beyond these, hybridization capture is widely applied in evolutionary biology and population genetics, museomics and ancient DNA recovery, forensic identification, pathogen surveillance, and environmental metagenomics, where high-quality data from defined loci is required.
Conclusion
Hybridization capture is the standard for scalable targeted sequencing because it can manage large panels and degraded sample types. While amplicon sequencing remains useful for small, specific variant sets, hybridization capture provides the uniformity and breadth required for more complex genomic projects.
Daicel Arbor Biosciences’ myBaits platform makes it easy to get started with hybridization capture-based NGS, a large collection of predesigned panels, with expert probe design support, scalable kit options, and compatibility with any NGS workflow. Learn more about our comprehensive targeted NGS hybridization capture kits.
References
- Wen-Hsing Cheng et al. (2017)Comparison of Capture-Based Targeted Next-Generation Sequencing vs. Amplicon-Based Targeted Next-Generation Sequencing for Mutation Profiling of Lymphoma. Journal of Molecular Diagnostics, 19(4), 508-519. https://www.jmdjournal.org/article/S1525-1578(17)30425-7/fulltext
- Samorodnitsky et al. (2015). Evaluation of Hybridization Capture Versus Amplicon-Based Methods for Whole-Exome Sequencing. Human Mutation, 36(9), 903-914. https://onlinelibrary.wiley.com/doi/10.1002/humu.22825
- Biocompare (n.d.) The Basics of Hybridization Capture. Biocompare White Paper. https://www.biocompare.com/Media/37/Document/BC_FL_NGS_TheBasicsOfHybridizationCapture.pdf



 Bluesky
Bluesky