Overview
By focusing sequencing efforts on specific regions of interest, targeted sequencing technology can increase the scale and efficiency of next-generation sequencing (NGS) research by orders of magnitude. Using myBaits hybridization capture kits in any NGS workflow can lower your sequencing costs by reducing irrelevant data from your analysis.
Cost savings—Focus your NGS on targets of interest, only pay for data you need.
Superior performance—Optimized chemistry and protocol for high, even coverage.
Probe design service—Includes project and panel design assistance from our scientists.
Open platform—Compatible with any NGS library preparation system.
Scalable—Different panel and kit sizes available for any project scale.
Complete solution—Convenient kits include hybridization and wash reagents.
myBaits kits are for research use only and are not validated for diagnostic or therapeutic purposes.
Technology
Enhance your sequencing efficiency with myBaits
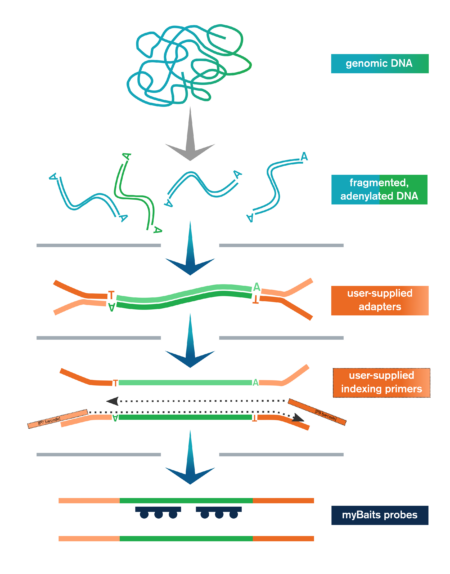
Daicel Arbor’s proprietary flexible and customizable oligo pool synthesis technology powers our myBaits hybridization capture kits. In-solution hybridization capture uses pools of thousands of custom biotinylated oligonucleotide molecules to selectively extract specific regions of interest in an NGS library prior to sequencing. This dramatically increases sequencing efficiency, minimizing irrelevant data from your analysis.
Pair with library prep for a complete workflow
The Library Prep Kit for myBaits provides reagents to prepare NGS libraries from dsDNA samples (with user-supplied adapters) as well as for the post-capture amp step of the myBaits protocol. Paired with myBaits, the Library Prep Kit offers a seamless solution to generating high-quality targeted DNA sequencing data.
myBaits kits are for research use only and are not validated for diagnostic or therapeutic purposes.
Applications
Target enrichment via hybridization-based capture is a powerful and versatile NGS tool. myBaits kits have been used successfully in thousands of research projects for a wide variety of applications, including:
- Variant/marker discovery
- Gene/exon/SNP re-sequencing
- Microbes / pathogens
- Metagenomics
- Species identification
- Genotyping
- Degraded and ancient DNA
- Phylogenetics and evolutionary biology
myBaits kits are for research use only and are not validated for diagnostic or therapeutic purposes.
Resources
Product Literature
Poster
A universal targeted sequencing system for any high-throughput sequencing platform
Application Note
myBaits kits are for research use only and are not validated for diagnostic or therapeutic purposes.
FAQs
No. myBaits hybridization capture kits have always been-- and will continue to be-- compatible with any user-provided upstream NGS library preparation system that is appropriate for a given project.
However, for your convenience, we are now offering our own powerful kit for preparing NGS libraries from most types of DNA samples, which is relevant for many myBaits hybridization capture projects. This new product (Library Preparation Kit for myBaits) is intended to be used for the preparation of NGS libraries from dsDNA samples prior to hybridization capture with myBaits.
In addition to the upstream library preparation steps, the Library Prep Kit for myBaits includes the reagents necessary for performing the post-capture amplification step of the myBaits protocol.
- Biotinylated RNA probes, with sequences corresponding to your custom design or a predesigned catalog option
- Hybridization and wash reagents
- For myBaits Custom kits, optional custom probe design informatics service (= expert bioinformaticians design and filter bait sequences and provide summary report and recommendations)
You will receive enough probes and reagents for performing the stated number of individual capture reactions of your kit size (e.g., 16 reactions) according to our current protocol. Please note that there are some additional reagents and equipment you will need to supply in order to perform a myBaits capture. Please review the list of required materials in the applicable myBaits manual to make sure you have everything you need before starting your experiments.
We also offer reagents for preparing libraries from DNA samples in advance of performing the myBaits hybridization capture step. Please visit Library Prep Kit for myBaits for more information.
If you are looking to outsource your project to a full-service laboratory and bioinformatics services group, please visit our myReads NGS laboratory and bioinformatics services page for more information about our comprehensive targeted sequencing service options (library preparation, target capture, next-generation sequencing, and optional analysis).
In this context, we use the terms interchangeably. Some fields prefer one term over the other, so we use both terms.
For new myBaits Custom baitset designs, the estimated manufacturing lead time is ~3-4 weeks minimum, starting from when your order is received and you have approved the final design. In addition, please consider that if you utilize our included bait design services, we will typically be in correspondence for an additional upfront period (up to several weeks) regarding a design before manufacturing can begin. Please also remember to accommodate any additional time for your collaborators to approve the final design, if applicable.
For myBaits Expert (catalog) kits or reorders of myBaits Custom kits with designs previously manufactured by Daicel Arbor Biosciences, the estimated manufacturing lead time is up to ~1-2 weeks from the time an order is received.
All myBaits kits include a specific protocol for their use as well as almost all of the reagents required to deploy them. In the manual, you will find the complete list of required supplies (reagents and equipment) that you will need in order to perform the captures.
Please see the applicable myBaits manual for detailed protocol instructions for enriching from Standard, High-Sensitivity, Long-Insert, or other specialty target/sample types.
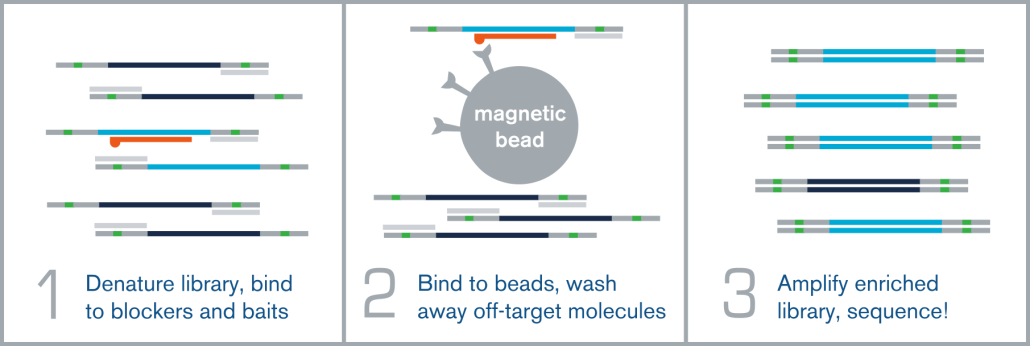
Hybridization capture is integrated into the overall NGS workflow immediately before sequencing on an NGS platform, such as Illumina. A fully sequenceable, barcoded/indexed NGS library (or pool of multiple libraries) is denatured, and allowed to anneal to complementary target-specific biotinylated probes/baits. These bait:library complexes are then bound to streptavidin-coated magnetic beads via the biotin on the probes, which are washed to remove non-specifically bound molecules. The remaining “enriched” library molecules are then released from the baits and amplified before sequencing.
Note! You may know the “hybridization capture” technique by another name, such as:
- Target enrichment
- Target capture
- Probe capture
- Exon capture
- Capture sequencing / sequence capture
- Hybridization sequencing / hyb-seq
- Hybridization capture / hyb-cap
Specific recommendations for per-library input mass for different enrichment project types can be found in the applicable myBaits manual.
Target capture necessarily requires subjecting your libraries to a bottleneck, wherein target molecules are captured and therefore enriched, and non-target molecules are therefore removed. To have sufficient unique molecules for good sequencing coverage of your targets, successful captures DEPEND on the input of sufficiently complex libraries.
For best results, it is recommended that only amplified (non-PCR-free) NGS libraries are used for target capture. This provides multiple copies of each starting template molecule, increasing the chance of each individual molecule getting enriched. However if you need more starting material to reach the recommended amount, it is generally preferable to generate more library from fresh genomic DNA or a new batch of indexed library, rather than through extra amplification. This is because while some amplification is good, over-amplification risks reducing the observable complexity of your libraries through the uneven action of PCR bias, as some molecules will become relatively more abundant while others become rare. This is also true for manipulating your libraries after capture: amplify your post-capture libraries the minimum number of cycles necessary to reach the molarity required by your sequencing facility.
The applicable myBaits manual covers some common technical questions and troubleshooting topics at the end of each protocol. Please read through the relevant section first as it may answer your question. If you still have an issue, please contact us via email at techsupport_at_arbor.daicel.com or reach out to your most recent contact person for assistance.
When ordering your myBaits kit, please indicate the sequencing library configuration you intend to enrich. The standard adapter blocking reagent provided with the kit (Block X) is compatible with Illumina® TruSeq®-style or Nextera®-style libraries with single 6-12 bp or dual 6-12 bp indexing. These options cover the vast majority of currently available commercial library preparation systems intended for sequencing on any Illumina platform.
For different adapter configurations than those described above, we recommend ordering Custom IDT® xGen® Blocking Oligos. At a concentration of 1 μg/μL, custom adapter-blocking oligos can be used in lieu of myBaits Block X.
If you are not certain, or later decide to change your library prep kit, please contact us so we can instruct you on how to obtain the correct blocking oligos.
Yes! Our expert myReads team provides a range of in-house NGS services, including library preparation, target capture with myBaits, high-throughput sequencing, and optional bioinformatics analysis. Visit the Sequencing Services page to learn more about our comprehensive laboratory and sequencing service options!
myBaits kits are for research use only and are not validated for diagnostic or therapeutic purposes.
Need assistance?
Our scientists are ready to help you select the right Daicel Arbor Biosciences product for your research needs.